Our paper entitled “Mesoscale volumetric light field (MesoLF) imaging of neuroactivity across cortical areas at 18 Hz” has been published in Nature Methods.
We present a modular, mesoscale light field (MesoLF) imaging hardware and software solution that allows recording from thousands of neurons within volumes of ⌀4 × 0.2 mm, located at up to 350 µm depth in the mouse cortex, at 18 volumes per second (Video 1). Using our optical design and computational approach we show recording of ~10,000 neurons across multiple cortical areas in mice using workstation-grade computing resources.
Video 1 | Animated perspective rendering of neuron positions and calcium activity recorded using MesoLF in mouse cortex. Field of view: ⌀4000 × 200 µm. Depth range: 0–200 µm. Recording frame rate: 18 Hz. Real-time recording duration: 405 s. Playback speed-up: 25×. Labelling construct: AAV9-TRE3-2xsomaGCaMP7f.
Information flow across mesoscale, i.e., multi-millimeter-sized regions of the mammalian cortex is a key feature of high-level cognition and is known to underlie complex behaviors. Yet, tracing this information flow in a volumetric fashion at a cellular resolution and high speed has remained challenging. This is primarily because most established neuronal activity imaging methods, such as two-photon microscopy, rely on time-consuming point-by-point scanning of an excitation beam focus to read out neuronal activity, as reported by the fluorescence rate of designer proteins known as genetically encoded calcium indicators (GECIs).
MesoLF is based on Light Field Microscopy (LFM), a 3D imaging technique that can capture fluorescence emission from GECIs in a parallel fashion from an entire volume in a single exposure of a camera and thus offers more favorable scalability to mesoscopic volumes while achieving neuron-level discrimination. LFM relies on computational deconvolution to infer the imaged volume from the recorded raw data images.
Our lab has established Light Field Microscopy in previous publications as a versatile neural recording technique. While initially only suitable for rather transparent samples, we have more recently also demonstrated a machine learning algorithm termed Seeded Iterative Demixing (SID) that extends the reach of LFM into more strongly scattering tissue such as the mammalian cortex.
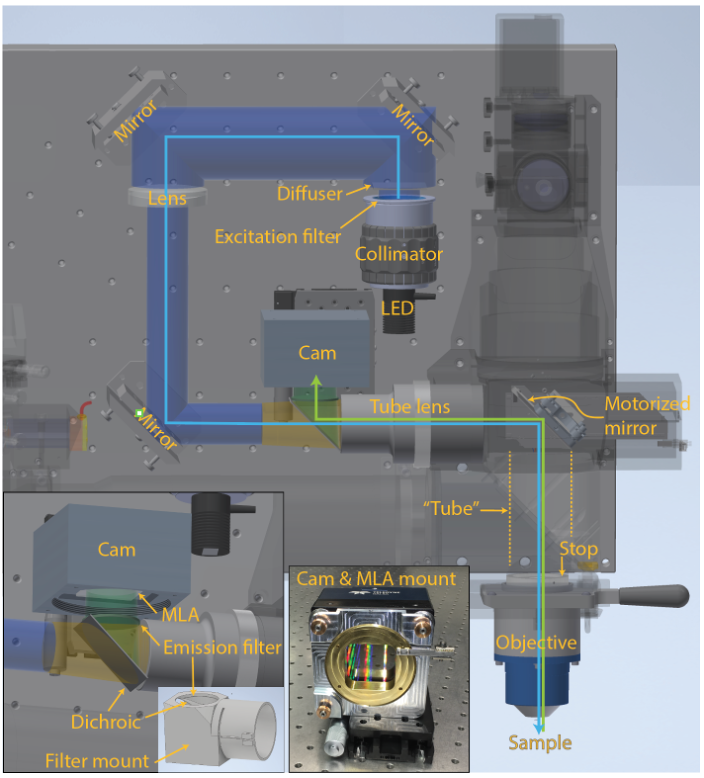
Figure 1 | MesoLF optical setup. Annotated rendering of main components of the MesoLF optical path. Components of the 2p-RAM system that are not relevant for the MesoLF optical path are shown semi-transparent for clarity. Left inset: Close-up of area around dichroic mirror and camera. Right inset: Photo of camera with custom microlens array (MLA) kinematic rotation mount, without MLA installed
In the present publication, we have for the first time realized mesoscopic light field recording solution, i.e., one that is capable of multi-millimeter-scale imaging in the mammalian cortex. Reaching this mesoscale has thus far been hampered by a lack of solutions for capturing mesoscopic fields of view at high optical resolution, and of computational tools that are suitable for tackling the resulting terabyte-scale raw data sizes. In addition to SID’s capability to extract signals in the presence of scattering, such computational tools must be able to address the unique challenges associated with faithful localization and extraction of neuronal signals at such scale, such as varying tissue morphology and non-rigid tissue deformation, while keeping computational cost at bay. We have designed the MesoLF optical system to be compatible with a widely used commercial mesoscopy platform known as the 2p-RAM system, which was originally conceived for multiphoton scanning microscopy but lacks well-corrected wide-field imaging capabilities. To implement MesoLF we designed and incorporated a custom tube lens in the optical detection path (Figure 1) and achieve diffraction-limited imaging of a ⌀4 mm field-of-view at NA 0.4 and 10× magnification.
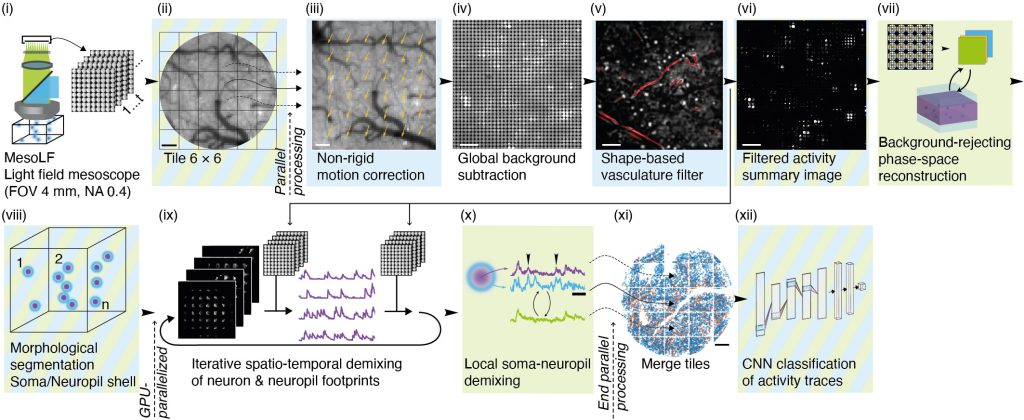
Figure 2 | Illustration of key steps of the MesoLF computational pipeline
The MesoLF computational pipeline (Figure 2, Video 2) is engineered from the ground up to maximize neuronal signal extraction performance at depth in scattering tissue at mesoscopic scale. The pipeline includes modules for correcting sample motion, background subtraction, masking out vasculature, rejecting fluorescence from above and below the imaged volume, and other preprocessing steps. It then performs computational reconstruction and signal demixing in an efficient, parallelized manner based on a novel variant of deconvolution algorithm. Special care has been taken to carefully reject contamination of neuronal activity signals from other neurons or neuronal processes in the local neighborhood surrounding each neuron. We perform quality control by using a convolutional neuronal network to reliably identify traces compatible with GCaMP transient characteristics with high signal-to-noise ratio (SNR).
Video 2 | Illustration of MesoLF pipeline processing a raw recording of calcium activity from mouse cortex, with key intermediate outputs. Panel in top right is comparison to established LFM reconstruction, showing that this method is incapable of undoing the blurring effects of light scattering. Bottom right panel shows MesoLF-detected neuron positions, with color indicating neuronal activity. Depth range 0–200 μm
We verified the performance of our MesoLF hard- and software solution by in vivo imaging in the cortex of mice expressing a modified version of the SomaGCaMP7f calcium indicator. In a representative example recording (Figure 3) we detected ~10,582 active neurons in the depth range of 0–200 µm, 8,076 active neurons in the depth range of 100–300 µm, and 4,746 active neurons in the range of 200–400 µm. The imaged volume contained all or the majority of the posterior parietal, primary somatosensory, primary visual, anteromedial visual, and retrosplenial cortical area (Figure 2, Video 1).
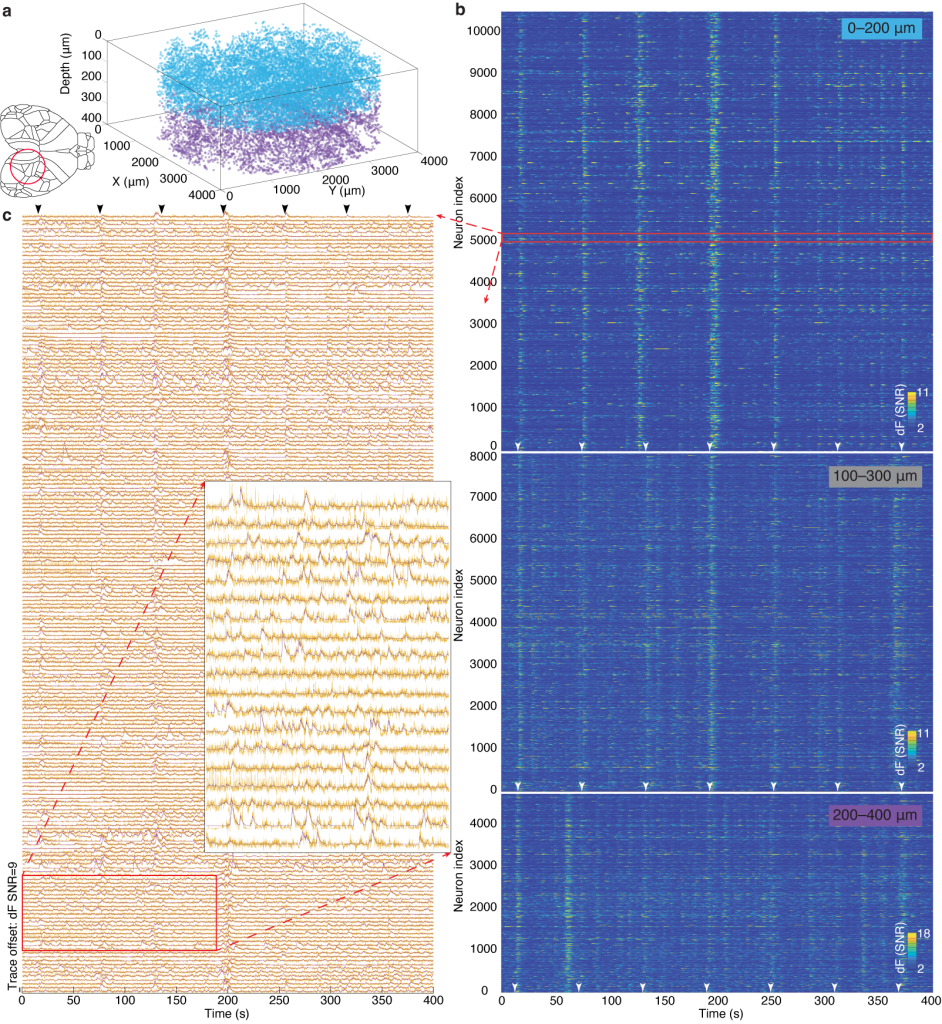
Figure 3 | MesoLF calcium imaging in the scattering rodent cortex.
(a) 3D rendering of single neuron positions within an overall volume of ⌀ 4 mm × 400 µm obtained by MesoLF from two subsequent recordings at 18 volumes per second in mouse cortex. Colors indicate recording depth range: blue, 0–200 μm; purple, 200–400 μm. Cortical areas contained in the imaged FOV include the posterior parietal, primary somatosensory, primary visual, anteromedial visual, and retrosplenial cortical area.
(b) Heat maps of denoised temporal signals extracted from three 405-second recordings at 18 Hz in mouse cortex at three different depth ranges (10,580 neurons in depth range 0–200 µm; 8,076 in 100–300 µm; 4,746 neurons in 200–400 µm.
(c) Stacked neuronal activity traces for region indicated by red rectangle in b. Yellow lines: un-denoised output of MesoLF pipeline. Violet lines: Denoising fit.
We carefully studied and quantitatively characterized the performance of our MesoLF method by comparing its output to verification recordings that were performed simultaneously using an established recording method (two-photon microscopy) and our new MesoLF solution, as well as by performing extensive simulations based on a realistic tissue model. We found good and robust performance across a depth range of 0–350 μm, as we describe in detail in the article.
Overall, our MesoLF solution delivers meso-scale high-speed optical recording of neuronal activity from up to 10,500 neurons within volumes of ⌀4 × 0.2 mm in the mouse cortex. This is made possible by exploiting the scalability inherent to LFM through a custom optical design, in combination with a set of machine learning innovations. The MesoLF optical and optomechanical design, as well as the software pipeline, will be available under an open-source license, thus lowering the entrance barrier to performing long-duration and high-throughput recording of volumetric calcium activity at mesoscopic FOVs.
Relevant publication:
Tobias Nöbauer*, Yuanlong Zhang*, Hyewon Kim & Alipasha Vaziri
Mesoscale volumetric light field (MesoLF) imaging of neuroactivity across cortical areas at 18 Hz.
Nature Methods (2023). doi: 10.1038/s41592-023-01789-z
https://www.nature.com/articles/s41592-023-01789-z