Our paper entitled “Video rate volumetric Ca2+ imaging across cortical layers using Seeded Iterative Demixing (SID) microscopy” has been published in Nature Methods.
We present a novel computational technique termed Seeded Iterative Demixing (SID) that allows for capturing neuronal dynamics in vivo within a volume of 900 × 900 × 260 µm, located as deep as 380 µm in the mouse cortex and hippocampus, and at a very high volume rate of 30 Hz. SID is based on Light Field Microscopy (LFM), a 3D imaging technique that our group has established in a previous publication as a versatile neural recording technique for weakly scattering specimen, such as larval zebrafish. SID extends LFM into more strongly scattering tissue such as the mammalian cortex by seeding a machine learning algorithm with remaining unscattered light and then iteratively “demixing” the effects of scattering in order to retrieve the neuronal activity signals from deep inside scattering tissue. The simplicity and scalability of LFM, coupled with the performance of SID opens up a range of new applications and is expected to propel its wide dissemination within the neuroscience community.
When light passes through a turbid medium such as the mammalian brain tissue, light rays are deflected (scattered), which leads to blurred images. This limits the accessible depths of most microscopy techniques to the most superficial layers of the brain when a wide-field configuration is used in which image voxels are captured in parallel.
The invention of scanning-based approaches such as two-photon microscopy has extended the reach of fluorescence microscopy to much greater depths this comes at the cost of temporal resolution.
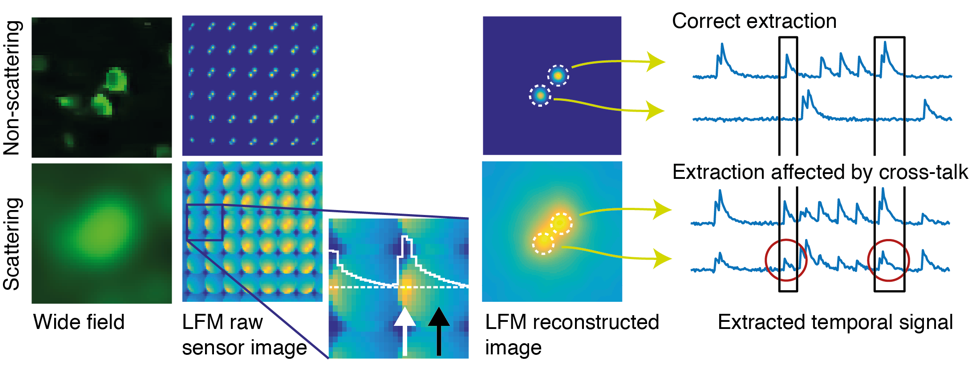
While we have previously conceived other strategies to speed-up the two-photon microscopy in this work, we pursue a different approach based on one-photon excitation and computational imaging to achieve fast, scan-less volumetric imaging of neuronal activity at extended depth: We record a sequence of frames from within scattering brain tissue in a light-field microscope, which in contrast to a conventional wide-field microscope not only records a spatial 2D image of the focal plane in the sample, but also the angle of incidence of the light rays that reach the camera sensor. This directional information not only allows for computational reconstruction of the sample volume from 2D camera images, but also provides an increased degree of robustness with respect to scattering. We devised an approach that makes use of this property, as well as the fact that the temporal signals from neurons labelled with fluorescent Calcium indicators vary in brightness as a function of neuronal activity.
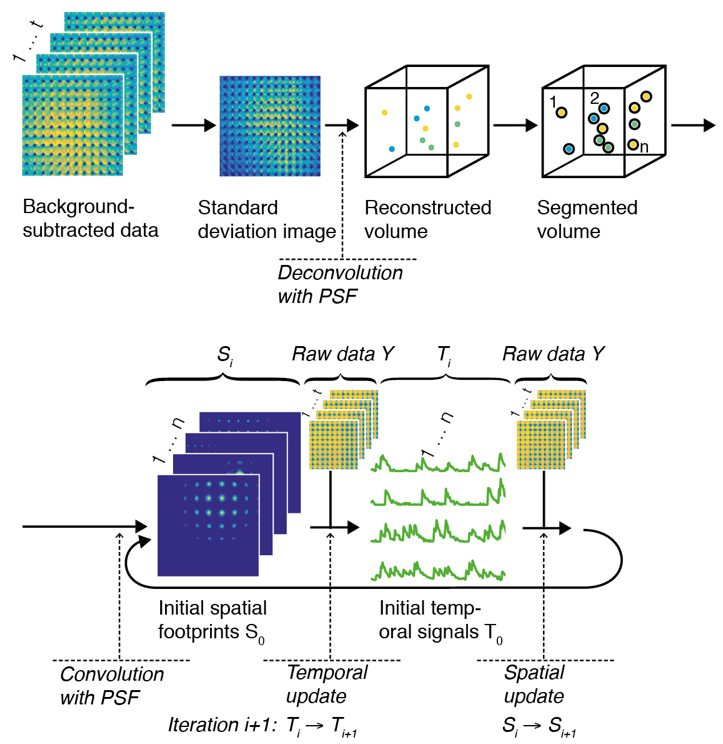
We start by exploiting the fact that at intermediate depths not all light is scattered, but some of it passes through the tissue un-deflected (“ballistically”). This may only be a small fraction of all light, but up to a certain depth enough to pick up in the raw data recorded by a light-field microscope, using statistical techniques. We then use these ballistic “neuron footprints” to seed a machine learning algorithm (a customized type of constrained non-negative matrix factorization), which then iteratively updates an estimate of the true neuron locations and their activity in time.
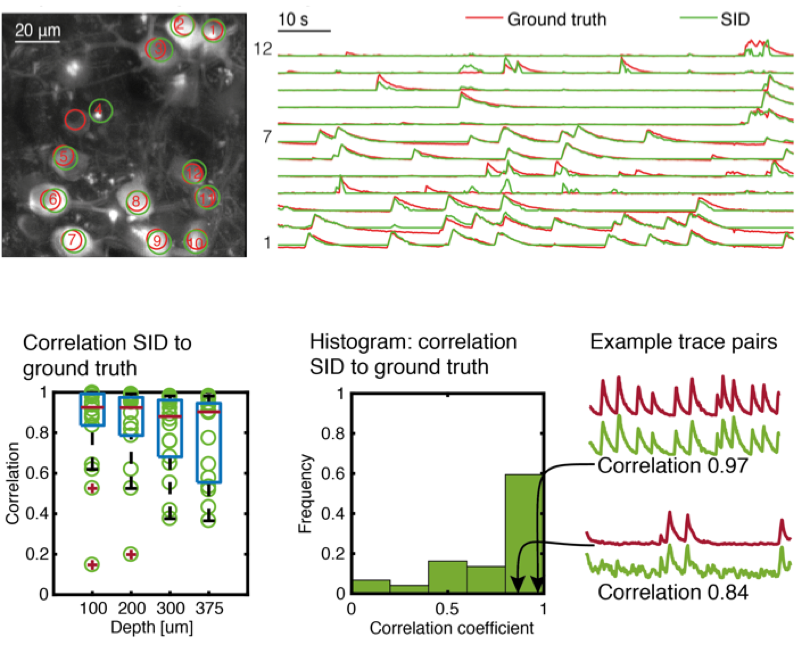
We demonstrated our method in mouse cortex and hippocampus and verified the quality of the extracted neuron positions and temporal signals by comparing it to ground truth two-photon microscopy data obtained in a custom-built, hybrid light-field/two-photon microscope.
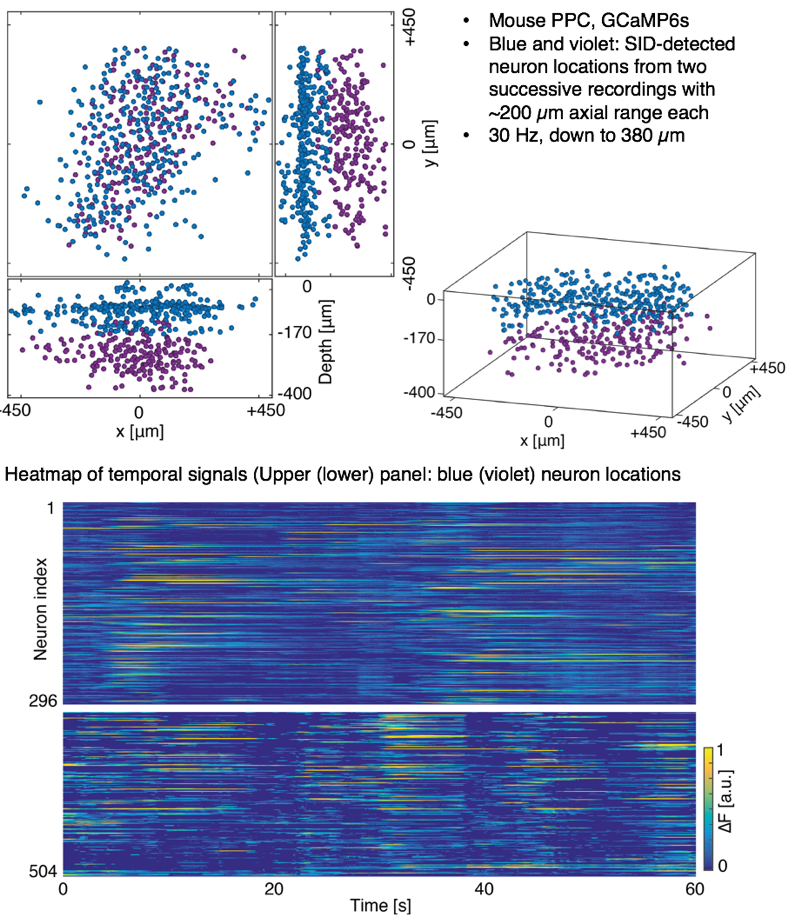
We demonstrate the performance of our SID microscope by recording the activity of hundreds of neurons in the cortex of awake behaving mice within a volume of ~900 × 900 × 260 located as deep as ~380 µm at a 30 Hz volume rate. While expectedly signal quality degrades at greater depths, in general we found good agreement with two-photon recordings for neurons spaced as closely as approx. 20 µm and up to a depth of 380 µm as quantified by the correlation coefficient between the SID-extracted and the ground truth data.
Moreover, a key advantage of our algorithm is that is allows for about 1000-fold reduction of the computational resources compared to the previous reconstruction-based approaches used for LFM. This paves the way towards integration of this approach with closed-loop optogenetics and behavioral paradigms. Overall, compared to other existing methods for high-speed volumetric Ca2+ imaging, SID stands out by its combined acquisition volume and speed, its simplicity and exceptionally low cost as well as its extreme scalability which we expect to contribute to its rapid dissemination.
Relevant publication:
Tobias Nöbauer, Oliver Skocek, Alejandro J Pernía-Andrade, Lukas Weilguny, Francisca Martínez Traub, Maxim I Molodtsov, and Alipasha Vaziri,
Video rate volumetric Ca2+ imaging across cortex using seeded iterative demixing (SID) microscopy
Nature Methods 14, 811-818 (2017).
(Download)
